Decoding Resistance: How Genomic Patterns Dictate Breast Cancer Treatment Outcomes
By NovaPress Editorial Board
Breast cancer remains one of the most significant health challenges globally, with advancements in treatment steadily improving patient outcomes. However, a persistent and formidable foe continues to vex oncologists and researchers alike: acquired therapy resistance. For many patients, initial successful treatments eventually lose their efficacy, leading to disease progression. A groundbreaking study published in the prestigious journal "Nature" sheds critical new light on this complex phenomenon, identifying specific genomic patterns that drive resistance and offering a roadmap for more effective, personalized interventions.
The Unseen Battle: Understanding Acquired Resistance
The fight against cancer is often a race against evolution. Cancer cells, under the selective pressure of therapeutic agents, can develop mechanisms to evade treatment. This acquired resistance is a major hurdle, transforming what might initially be a manageable disease into a relentless one. The "Nature" study, drawing on an expansive cohort of 6,927 tumour samples from 5,881 breast cancer patients, represents a monumental effort to dissect the genetic underpinnings of this resistance.
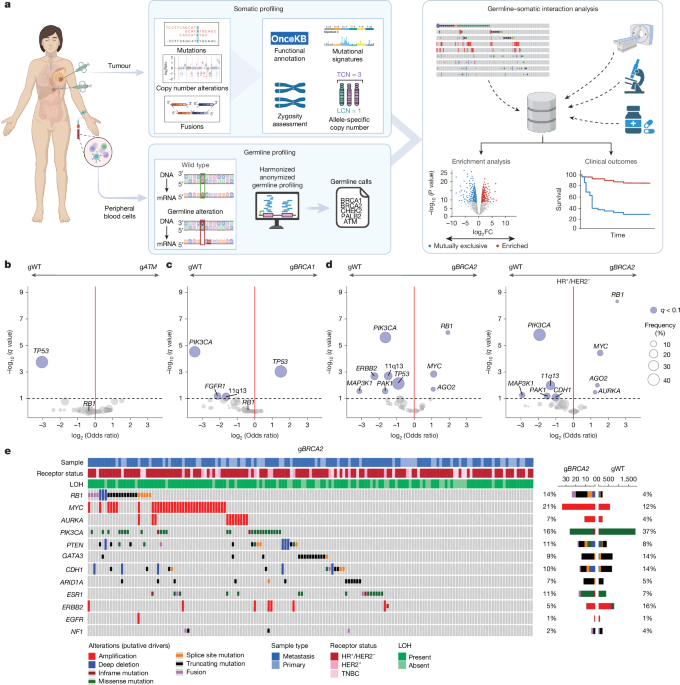
By performing prospective clinical tumour and normal DNA sequencing, researchers were able to meticulously map the genomic landscape of both the patient's inherent genetic makeup (germline) and the specific alterations within the tumor itself (somatic). This comprehensive approach allowed for an unprecedented view into the dynamic interplay between these genetic factors and their role in dictating treatment response.
Homologous Recombination Deficiency and Hemizygosity: Key Drivers Unmasked
At the heart of the study's findings are two critical genomic vulnerabilities: Homologous Recombination Deficiency (HRD) and hemizygosity. These terms, while complex, represent fundamental mechanisms through which cancer cells can become impervious to standard therapies:
- Homologous Recombination Deficiency (HRD): HRD refers to a cell's impaired ability to repair double-strand breaks in DNA, a crucial process for maintaining genomic integrity. While HRD can make cancer cells vulnerable to certain targeted therapies (like PARP inhibitors), the study highlights how its specific patterns, particularly when combined with other genetic events, can paradoxically drive resistance to other forms of treatment. Understanding the nuances of HRD status is therefore paramount.
- Hemizygosity: This refers to the state where only one copy of a gene or chromosomal segment is present, instead of the usual two. When key genes involved in drug metabolism or response become hemizygous, it can profoundly alter how a tumor reacts to therapy. The study reveals that specific patterns of hemizygosity, often arising from large-scale genomic rearrangements, are intimately linked with the development of resistance.
The novelty lies in identifying how the intricate interactions between germline predispositions (e.g., inherited mutations) and somatic alterations acquired during tumor development collectively define these "actionable genomic patterns." This means that it's not just one mutation in isolation, but the unique combination and context of these genetic events, that predicts how a tumor will respond to treatment, and crucially, how it might develop resistance.
Implications for Precision Oncology and Future Treatments
The findings from this Nature study are nothing short of transformative for the field of precision oncology. By pinpointing the specific genomic signatures that drive therapy resistance, researchers and clinicians gain powerful new tools:
- Predictive Biomarkers: The identified genomic patterns can serve as robust biomarkers, allowing clinicians to predict which patients are most likely to develop resistance to certain therapies. This enables proactive treatment adjustments, potentially before resistance fully manifests.
- Tailored Therapies: Understanding the underlying genomic mechanisms of resistance opens doors for developing highly targeted therapies that counteract these specific vulnerabilities. If a tumor develops resistance via a particular HRD pathway, new drugs could be designed to exploit that altered pathway.
- Drug Repurposing: The insights might also guide the repurposing of existing drugs, identifying combinations that could overcome resistance mechanisms identified by these genomic patterns.
- Enhanced Clinical Trial Design: Future clinical trials can be designed to stratify patients based on these genomic patterns, leading to more efficient and informative studies that identify effective treatments for specific patient subsets.
A New Era in Breast Cancer Management
This research underscores the power of large-scale genomic sequencing and sophisticated analytical approaches in unraveling the complexities of cancer. By moving beyond a one-size-fits-all approach, and instead focusing on the unique genomic fingerprint of each tumor and patient, we are entering a new era of breast cancer management.
The journey from groundbreaking discovery to clinical implementation is long, but studies like this provide the foundational knowledge necessary to get there. For patients battling breast cancer, this research offers a renewed sense of hope—the hope that through deeper genomic understanding, the challenge of therapy resistance can ultimately be overcome, paving the way for more durable and personalized cures.